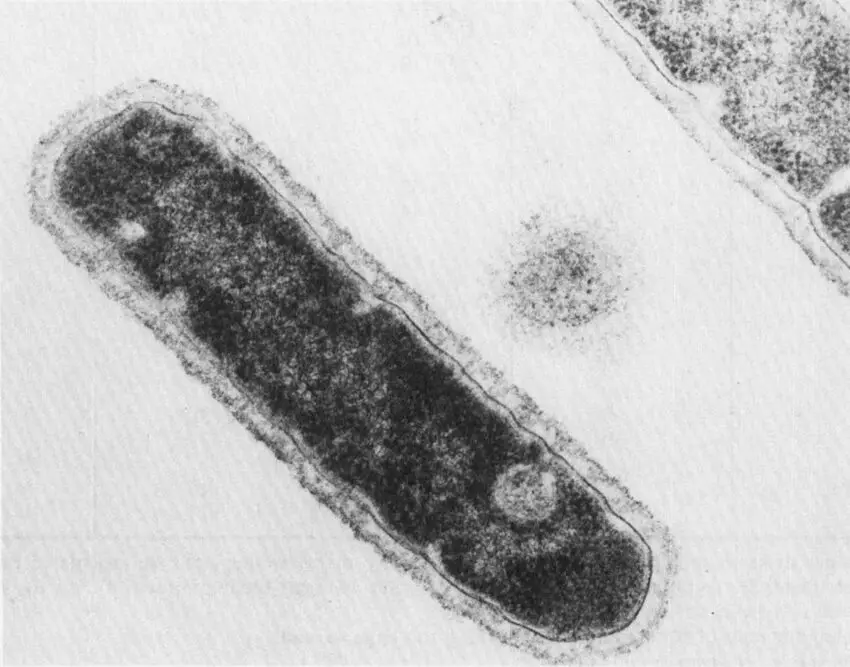
In 1966, microbiologist Thomas D. Brock collected bacteria from scalding hot springs in Yellowstone — environments previously thought hostile to all life.
The organism, Thermus aquaticus, thrives at 70–80°C. Its DNA polymerase — Taq polymerase — remains active at the extreme temperatures required for PCR thermocycling.
Kary Mullis conceived the Polymerase Chain Reaction in 1983 while driving along a moonlit California highway. The idea earned him the Nobel Prize a decade later.
DNA is a double-stranded molecule built from four nucleotide bases: Adenine (A), Thymine (T), Guanine (G), and Cytosine (C).
The two strands are held by hydrogen bonds between complementary bases: A pairs with T (2 H-bonds) and G pairs with C (3 H-bonds). GC pairs are stronger — this affects Tm.
The strands run antiparallel (5'→3' and 3'→5'). PCR exploits this to produce two identical daughter duplexes from every template molecule.
A thermocycler precisely controls temperature to drive three sequential molecular events. Each complete cycle takes 1–3 minutes.
Taq polymerase survives all three temperatures — unlike early PCR which required adding fresh polymerase every cycle. Yellowstone made full automation possible.
30 sec
30 sec
60 sec/kb
At high temperature, thermal energy overwhelms the hydrogen bonds holding the base pairs together. The double helix "melts" into two single-stranded templates.
Each strand now serves as a blueprint for copying. Taq polymerase tolerates 94°C unharmed — this is the key contribution of T. aquaticus to PCR.
GC-rich regions require higher temperatures (3 H-bonds vs 2 for AT). An initial hot start at 95°C for 5 min fully denatures long templates.
As temperature drops, short synthetic oligonucleotides — primers (18–25 bp) — bind to complementary sequences on each template strand by Watson-Crick base pairing.
The forward primer binds the antisense strand, defining the 5' end of the amplicon. The reverse primer binds the sense strand on the opposite end.
Primer Tm, GC content (45–65%), secondary structures, and genomic specificity are all critical design parameters. Annealing T° is typically Tm − 5°C.
Taq DNA Polymerase — optimally active at 72°C — binds at the 3' end of each primer and begins synthesising a new complementary strand in the 5'→3' direction.
Free deoxyribonucleotide triphosphates (dNTPs) are incorporated base by base. Each NTP releases pyrophosphate, driving the reaction forward thermodynamically.
After extension, every original double strand has produced two daughter duplexes. One PCR cycle is complete — copy number has doubled.
Starting with 64 copies (2⁶) shifts the curve left — same shape, earlier Ct. After Ct, reagents deplete and real growth plateaus.
Standard PCR requires a DNA template. But critical pathogens — SARS-CoV-2, Dengue virus, Influenza — carry RNA genomes. Gene expression studies also target mRNA.
RT-PCR adds a first step: reverse transcriptase enzyme converts single-stranded RNA into complementary DNA (cDNA). This cDNA then enters standard PCR thermocycling.
One-step RT-PCR runs both in a single tube. Two-step separates them for flexibility. Either way, the PCR cycles you already know follow directly.
In TaqMan qPCR, each primer is paired with an oligonucleotide probe that also anneals within the amplicon region.
The probe carries two molecules: a Reporter dye (R, 5' end) capable of emitting fluorescence, and a Quencher (Q, 3' end) that absorbs that emission while the probe is intact.
When cool, primers and probes co-anneal. The reporter and quencher remain close — no signal is emitted yet. Energy transfer (FRET) between them keeps the fluorescence suppressed.
As temperature rises, Taq polymerase binds at each primer's 3' end and begins extending toward the probe.
Taq's 5'→3' exonuclease activity physically degrades the probe as it encounters it — separating reporter from quencher. The probe is destroyed in the process.
Once free, the reporter emits fluorescence that accumulates with every cycle. This signal is proportional to the amount of amplicon produced — detected in real time.
In qPCR, a fluorescent signal accumulates in proportion to amplicon quantity each cycle — no gel required. The reaction is monitored in real time.
SYBR Green intercalates into any dsDNA. TaqMan probes provide sequence-specific signal via 5'→3' exonuclease activity of Taq — cleaving a quencher-dye probe as it extends.
The Ct value (Cycle Threshold) marks where fluorescence crosses the threshold. Lower Ct = more starting template. Every ~3.3 cycles = 10-fold difference in starting quantity.